 
Where p 0 and q 0 are the initial frequencies of the two alleles at a locus. Var (p) = after one generation of genetic drift for diploid organisms.Īfter many generations of genetic drift, an equilibrium will be reached. 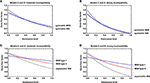
The expected variance in the frequency of an allele (call this frequency p) subject to genetic drift is: 
We can calculate how much genetic drift we expect to find in a population if we know the effective population size. An analysis of field populations of the wheat pathogen Mycosphaerella graminicola indicated N e of at least 70 strains per square meter (Zhan et al., 2001). N e is not easy to define for fungal pathogens that undergo a mixture of sexual and asexual reproduction because the absolute number of individuals can be very large while the number of different genotypes that sexually recombine can be relatively small. N e is not easy to quantify because it is affected by reproduction and breeding strategies (inbreeding, outcrossing, asexual reproduction), and is dependent on the geographical area over which a population is sampled. N e can also be thought of as the number of genetically distinct interbreeding individuals in a population. N e is a theoretical number that represents the number of genetically distinct individuals that contribute gametes to the next generation. 
N e is rarely the actual number of individuals in the population (also called N or the census size). The magnitude of genetic drift depends on N e, the effective population size, for the population. A founder event occurs when one or two infected plants slip through a quarantine and introduce a disease into an area where the disease did not previously exist. A founder effect occurs when a small number of individuals, representing only a small fraction of the total genetic variation in a species, starts a new population.periods of hot, dry weather or a deep freeze). harvest of the crop), or when the environment changes to prevent infection of the plant or to kill the pathogen directly (e.g. A genetic bottleneck, or severe reduction in population size, occurs when the plant population is removed (e.g.Small recurring population size occurs when there are not many host plants in the area to infect, or when the environment is not optimal for infection.We will consider these in the context of pathogen populations in plant pathosystems: This may be especially important in natural ecosystems where both plants and pathogens are likely to have a patchy distribution where each patch is a small population.īecause allele frequencies do not change in any predetermined direction in this process, we also call genetic drift "random drift" or "random genetic drift." The sampling error can occur in at least three ways. Drift is probably common in populations that undergo regular cycles of extinction and recolonization. Drift increases the inbreeding coefficient and increases homozygosity as a result of removing alleles. Genetic drift leads to fixation of alleles or genotypes in populations. Random drift is caused by recurring small population sizes, severe reductions in population size called "bottlenecks" and founder events where a new population starts from a small number of individuals. Genetic drift is a random process that can lead to large changes in populations over a short period of time. Genetic drift is a process in which allele frequencies within a population change by chance alone as a result of sampling error from generation to generation. But small population sizes also introduce a random element called genetic drift into the population genetics of organisms. It should now be clear that population size will affect the number of alleles present in a population. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
